Within the framework of the ROSiE project, a number of workshops took place examining the challenges related to research ethics, research integrity and the FAIR principles arising for Open Science Infrastructures (OSI). Participants formulated several potential solutions to address the challenges previously identified and allow responsible OSI. Potential solutions are concrete mechanisms to develop, and often rely on real-life experience and examples from participants’ practices.
- Building Integrated Systems that allow harmonization of data storage, organisation and evaluation of quality. This may especially help to address re-use and Interoperability challenges.
- Creating Common Charters: the creation of a charter to be adopted by all the actors collecting and sharing the data into a same infrastructure or platform[1]. This common charter may help address the challenges mentioned above.
- Implementing Data Justice Plan as worded by a participant:
“One of the things that we’ve come up with in that work is that, rather than just having data management plans which look at the data life cycle throughout the project, that are mostly focused internally, we should supplement those with something that we call a data justice plan which would allow for projects to really showcase, and then track how the data benefits the citizens and what sort of citizens contribute. And what is the benefit of our data for the community, for the citizens / for the participants, and have that documented, alongside data management, could be part of a data management plan.”[2]
This echoes the CARE principles previously mentioned and may help address the digital divide challenge and the reuse challenge, among others.
- Creating Playbooks: short and easy playbooks to help understand how to create a DMP[3].
- Rewording Consent Forms to adapt them to the context of OS, thus addressing the challenge of Informed Consent[4].
- Implementing an OSI Service Level Agreement to be adopted by platforms, that describe expectations, procedures and guidelines of the OSI and the data gathered – for example, related to the FAIR principles or on how platforms will work together. Such agreement may help address all the challenges presented above.
- Developing Wikidata Models refers to the development of an expertise community annotation and curation process to address data quality & integrity and ensure trust.
“And in one, one way we’re also looking into kind of the wikidata model […], where you could go and edit data or upload data. So in particular, we are thinking about this as a community curation and community annotation model. […] So we want to bring that sort of, you know, community level where different expertise can come in and annotate or provide input on the quality of the data.”
This goes in combination with a Flag and Annotation System (i.e., to identify errors, missing info etc.) which can lead to a “data quality score or trust score”. This model can work with the implementation of a Persistent Expertise Profile (a guarantee of the credibility of the user and therefore of the quality of the flag and annotations it generates) that allows the origin of each piece of data, annotation or flag to be informed.
“And of course we also try to show the value. So for example, we spend a lot of time on thinking about persistent identifiers and metadata and repositories so we can show the researchers, “okay, if your data is in a trusted repository with a persistent identifier that’s linked to your publication, you know, others can find it, others can cite it”. So then you can you can sort of think about the value of it. So sort of the carrot.”
- The persistent expertise profile can also help meet the challenge of contribution recognition.
- Lowing Data “Quality”: some offered to use lower quality data for better inclusivity (addressing thus the Digital Divide challenge). For example, it may involve the use of XML-TEI[5] encoding or the use of platforms that allow the adjustment of video bandwidth according to the quality of the users’ connection.
In conclusion, the main challenges mentioned by the participants to the workshops can be summarized by this visual representation:
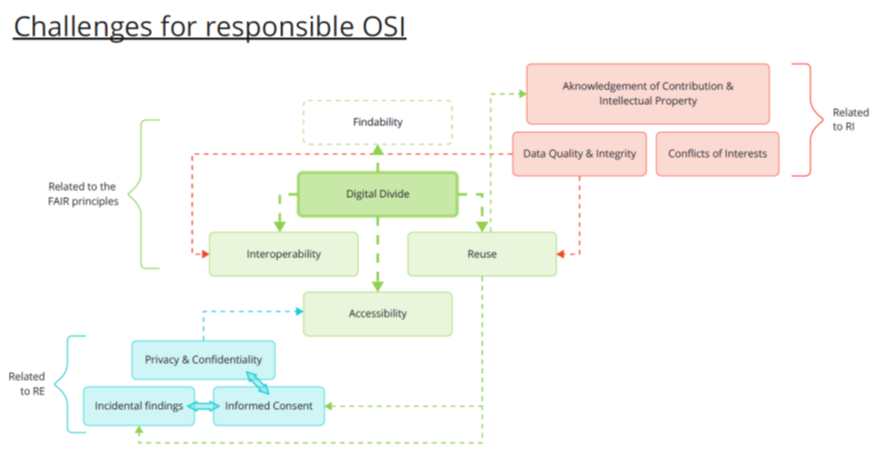
[1] See for example : https://rdmkit.elixir-europe.org/
[2] All quotes provide from the discussions in the workshops.
[3] See for example : https://cetaf.org/resources/best-practices/
[4] See for example : https://elixir-europe.org/about-us/how-funded/eu-projects/converge
[5] For more information, see: https://tei-c.org/
This passage is part of D6.2: Final analysis and mapping of existing European and national OS infrastructures with regard to promoting responsible OS written by Carole Chapin, Nathalie Voarino.
